Projects
You can learn more about ongoing specific projects below.
Featured
 Explainability + Genomics
Explainability + GenomicsDimensionality Reduction Algorithms with a focus on explainability for single cell transcriptomics experiments.
More
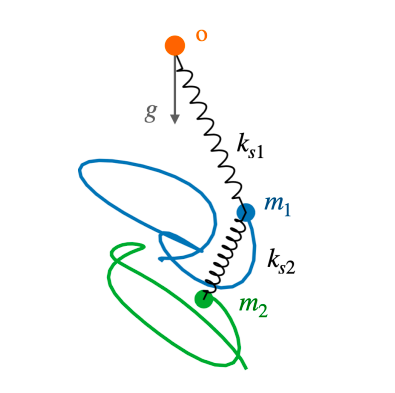 Physics + ML
Physics + MLPhysics informed machine learning tools for understanding self-organizing systems. Multi-agent simulations, graph neural networks and symbolic regression models for in silico active matter, developmental patterns and ecology.
 In vitro vs in vivo systems
In vitro vs in vivo systemsTransfer learning and multi-modal learning for organoid to scRNA-seq integration and comparison. What gene regulatory signatures encountered in vivo are recapitulated in organoid cultures?
 Mechanics + Transcriptomics
Mechanics + TranscriptomicsBayesian machine learning methods for inferring mechanical properties and morphological features of cellular aggregates with the goal of quantifying their interaction with transcriptomics.
 Infer + Perturb
Infer + PerturbExperimental design and multi arm bandits for optimal perturbations in transcriptomic studies.
